Tissue immunity, decoded
We use spatial transcriptomics, single-cell sequencing, and genetic perturbation to understand the cellular logic of tissue-resident immune responses — with therapeutic implications for infection, autoimmunity, and cancer.
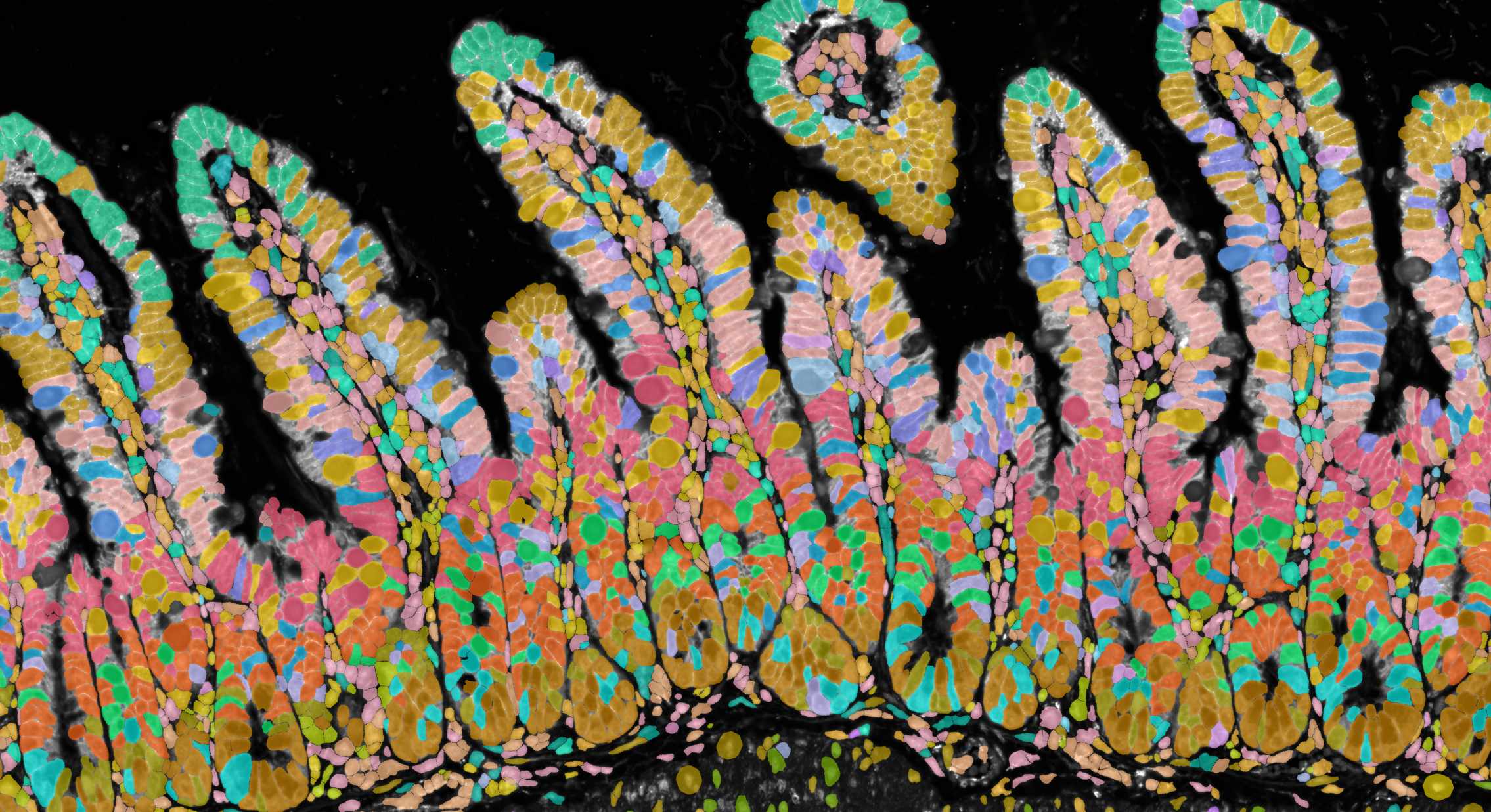
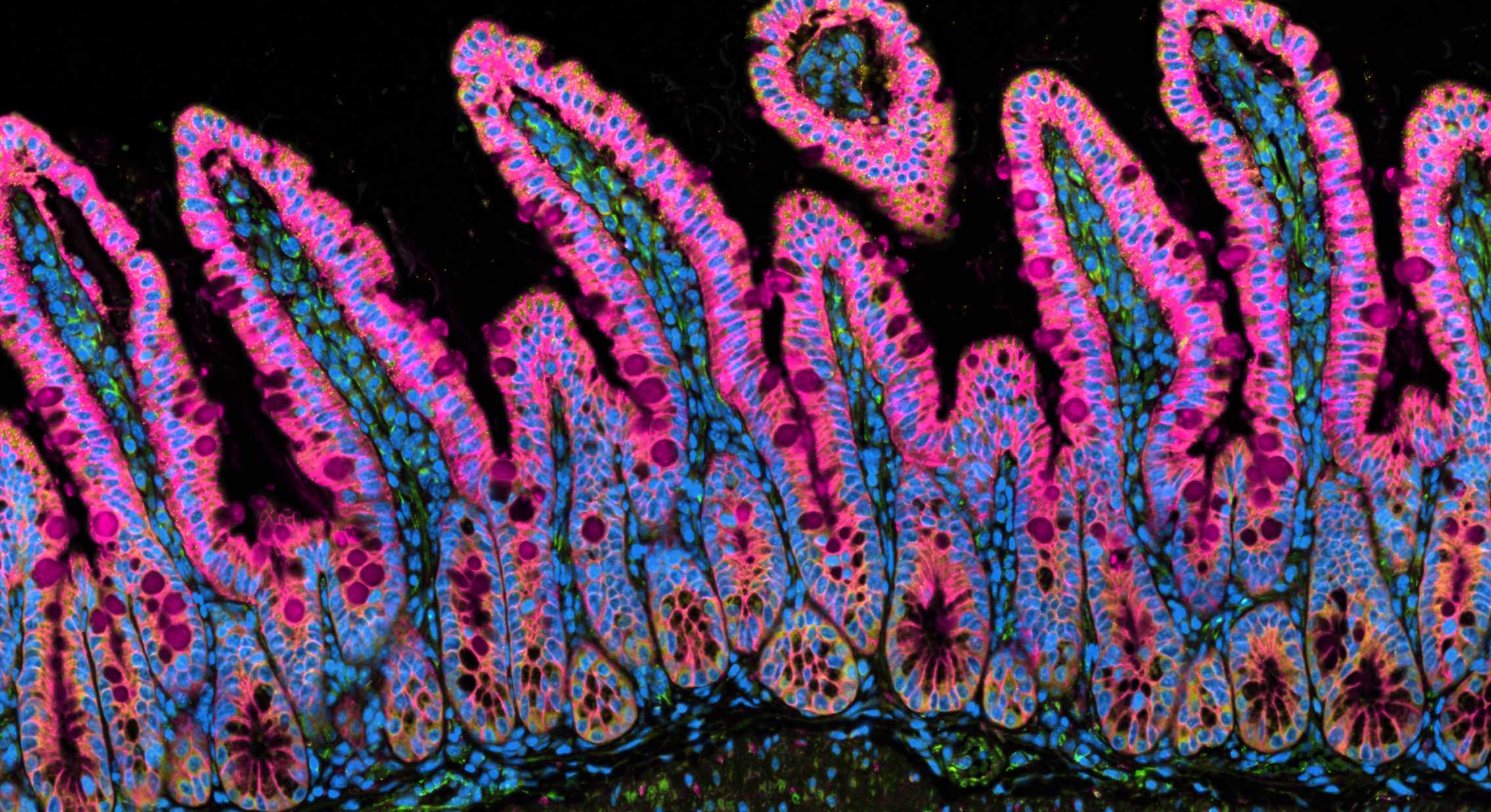
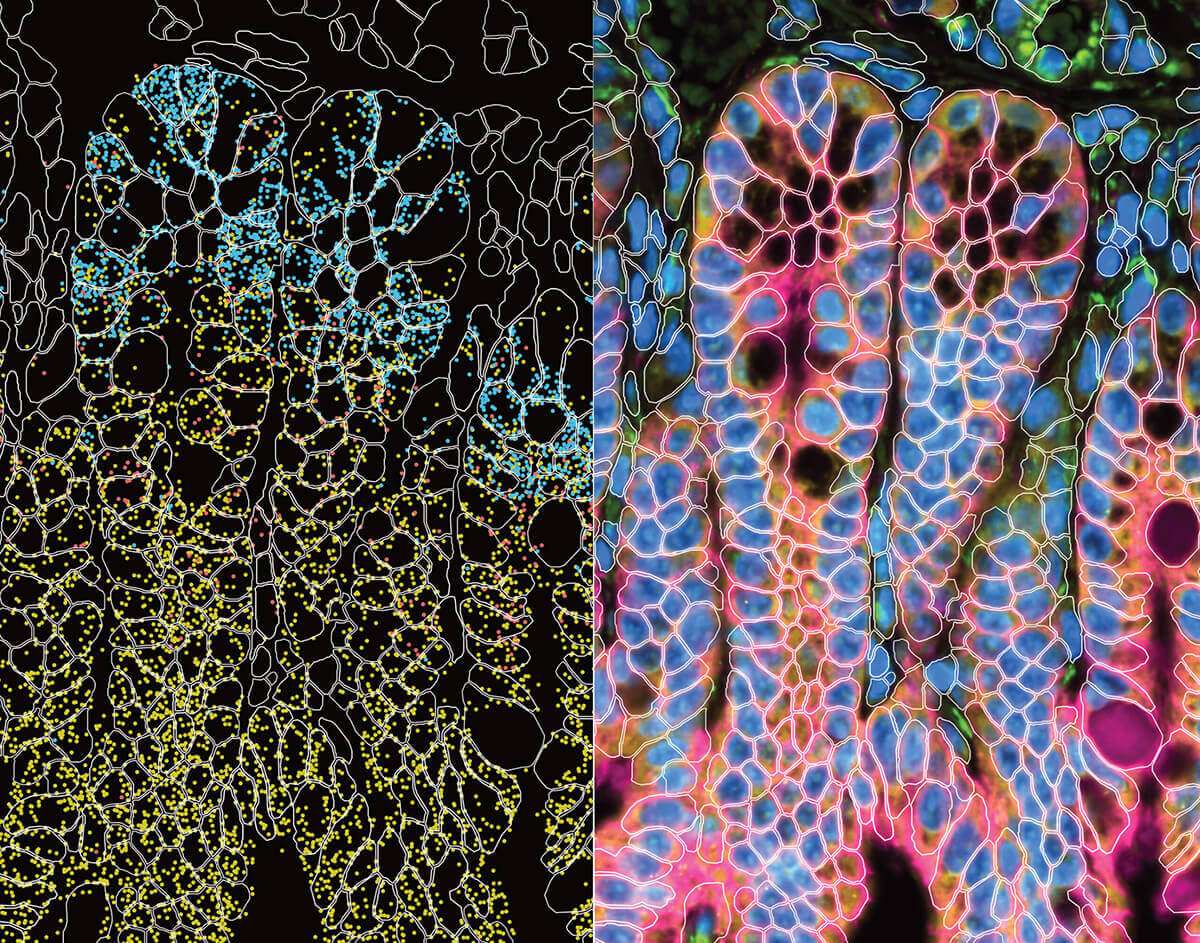
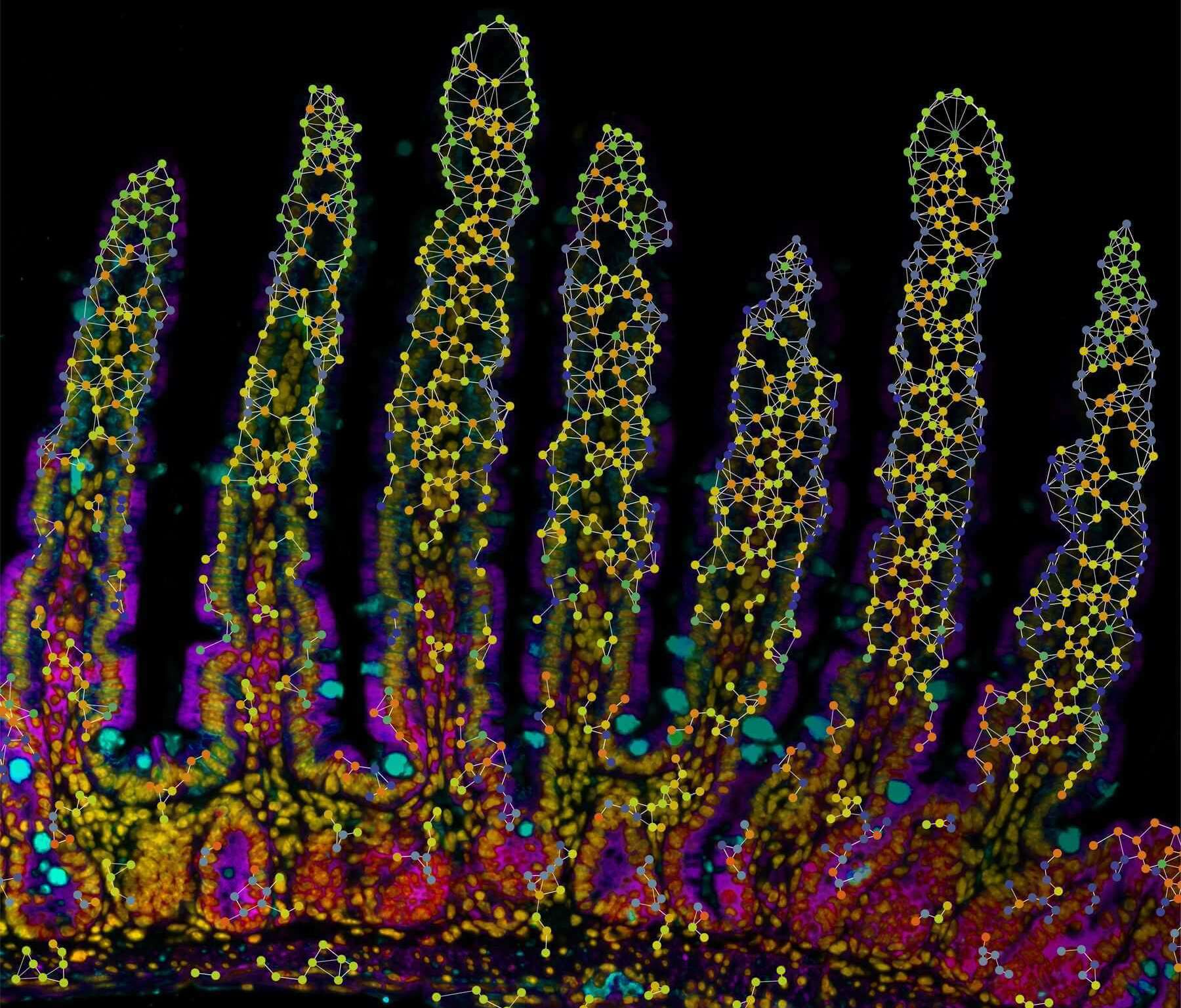
Xenium experiment of a mouse small intestine. Immunofluorescence and cell outlines are shown.
A selection of current projects is highlighted below. For a full overview of ongoing and past work, visit the projects page.

Spatial Orchestration of small intestinal tissue-resident T cells
Using spatial transcriptomics, we try to overcome limiations of single-cell sequencing and study T cells in intact tissues to understand which cell-cell interactions, gradients and cellular niches promote memory formation in barrier tissues.
View project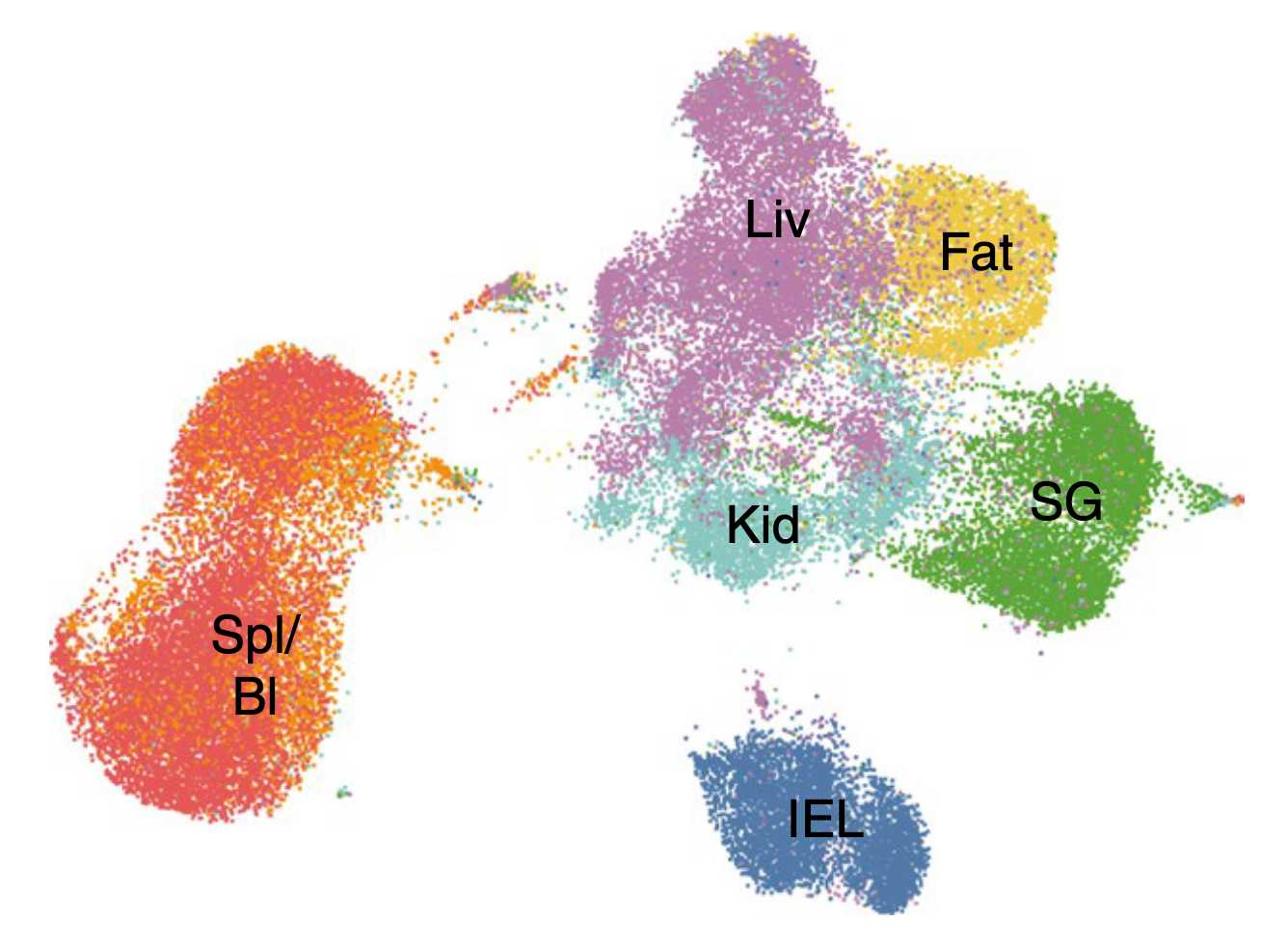
Transcriptional regulation of tissue-resident memory cells
How do tissue-resident memory cells adapt to unique tissue microenvironments? How do they sense environmental signals? How are they incorporated? Using mouse models of acute viral infection, combined with genetic perturbations and single-cell sequencing, we explore the transcriptional networks that govern the acclimatization of T cells to various barrier tissues.
View project